|
9/28/2023 0 Comments Tree of life biologyTrees inferred from a small set of universal core genes have typically supported a long branch separating the archaeal and bacterial domains. PLOS Biology, 14(12): e2001624.Core gene phylogenies provide a window into early evolution, but different gene sets and analytical methods have yielded substantially different views of the tree of life. Lifemap: Exploring the Entire Tree of Life. Nothing in Biology Makes Sense Except in the Light of Evolution. I hope that developmental biology will be a field for which information could be displayed on Lifemap as well? I will be happy to consider such options if some interest is declared!ĭobzhansky, T. These examples show how flexible Lifemap is at welcoming information from various sources. I also added links on every node and tip to the corresponding web page on the NCBI web site and on other taxonomy web resources. It is useful for easily identifying species, close to our species of interest, whose genome is available. This information can be displayed directly on the interactive tree. Because I personally work on comparative genomics, I started adding information on the number of genomes that are fully sequenced according to the NCBI database (Figure 1, bottom-left). In the near future, I would like the NCBI and the OTOL versions of Lifemap to be proper “hubs” giving access to a multitude of information of relevance to researchers in various fields. 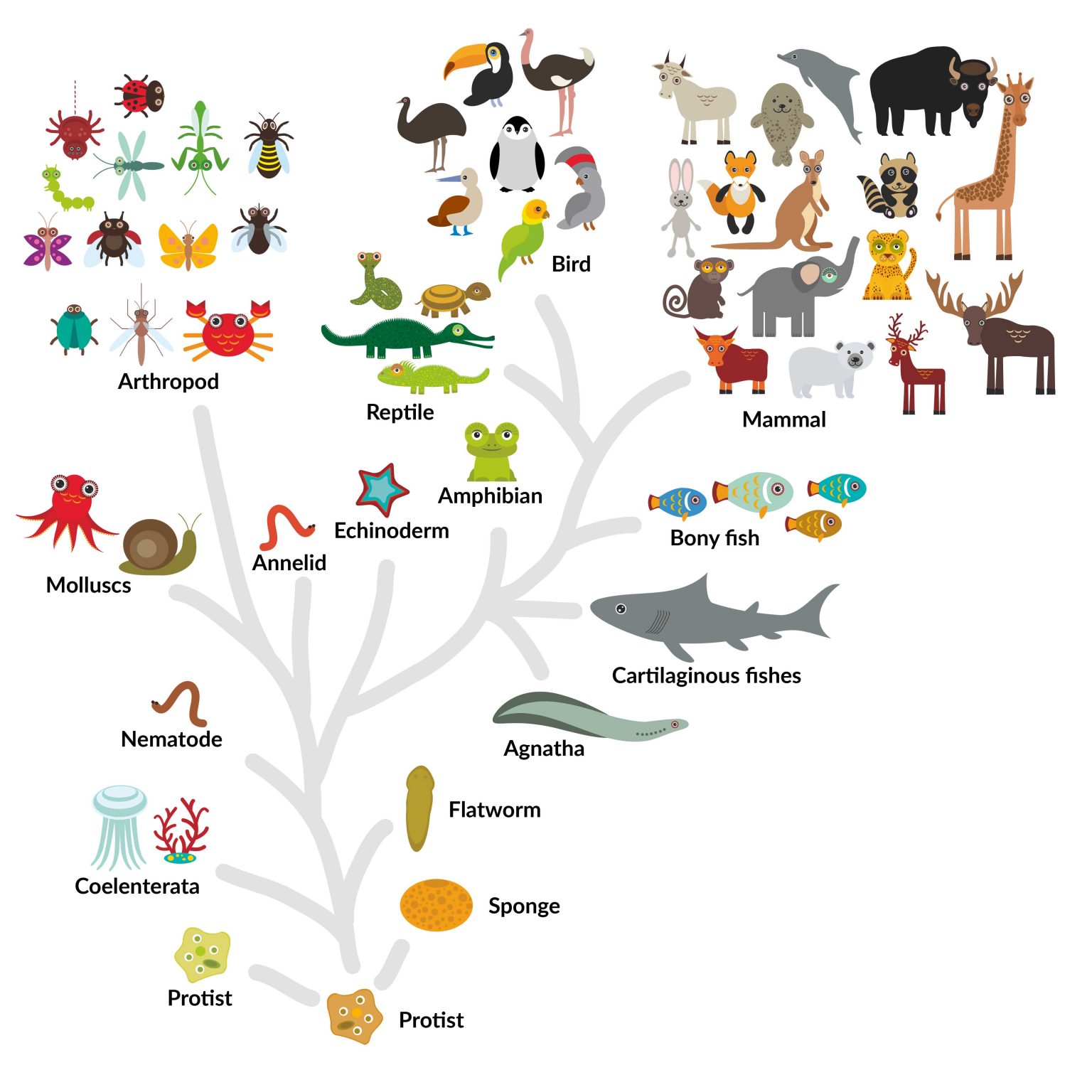
A first version displays the tree proposed by the Open Tree of Life ( OTOL) project, a second one displays the whole NCBI taxonomy, and a third one displays a simplified version of the taxonomy with pictures and descriptions when clicking on the nodes (this last version is also available as an App for phones and tablets on the Android market). Lifemap comes in three versions, depending on the tree that is displayed and the information that is associated to nodes and tips when clicking. Top-left: the three domains of the Tree of Life Top-right: inside Bacteria Bottom-left: orange bubbles represent the number of genomes fully sequenced Bottom-right: a “route” from Homo sapiens to Mus musculus. Finally, an efficient search engine allows for an immediate retrieval of any node or tip by its Latin or its common name.įigure 1. It is also possible to click on every node and tip of the tree to have access to additional information (description, picture, link to external resources, etc.), and to search for “routes” in the tree of Life (Figure 1). The exploration of the ToL in Lifemap is similar to the exploration of a geographic map: by zooming and panning. Lifemap relies on tools that were developed for cartography, especially in the context of the OpenStreetMap project, and proposes a new way of representing hierarchical structures. This tool solves many issues that previous methods had, such as the possibility to display nodes with more than two descendants (multifurcations) and the possibility to explore smoothly very large trees of more than 2 million species. In a paper published last December in PLOS Biology ( de Vienne, 2016), I described an online tool, called Lifemap, for exploring the entire Tree of Life (or any large tree) in a zoomable interface (Figure 1). 
But displaying this information in a clear and simple way that is also convenient and useful to biologists outside the field of evolutionary biology has been largely underexplored. This implies better species sampling, more sequencing, improved evolutionary models and new algorithms for phylogenetic tree reconstruction. Many researchers in evolutionary biology are working on improving the quality and the comprehensiveness of this ToL. 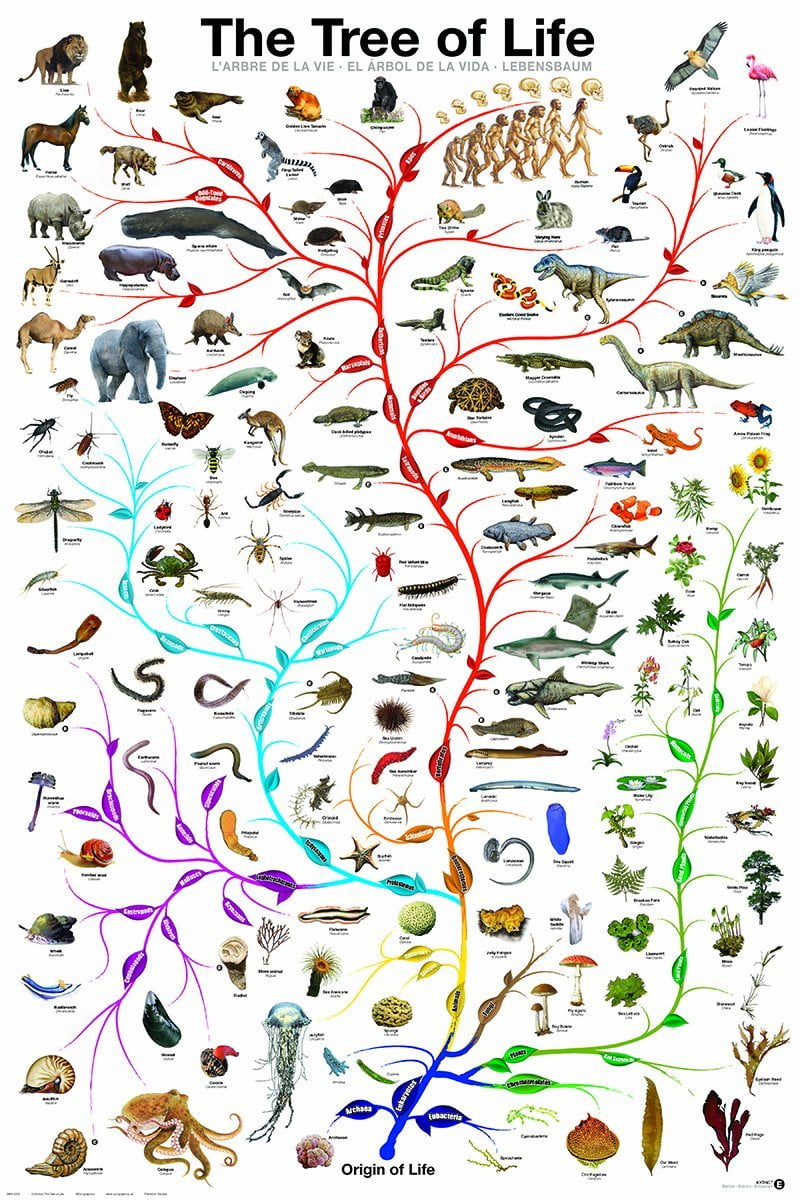
Our knowledge of the evolutionary relationships between all known organisms, the so-called Tree of Life (ToL), is crucial in all fields of biology. “Nothing in biology makes sense except in the light of evolution” (Dobzansky, 1973).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
